 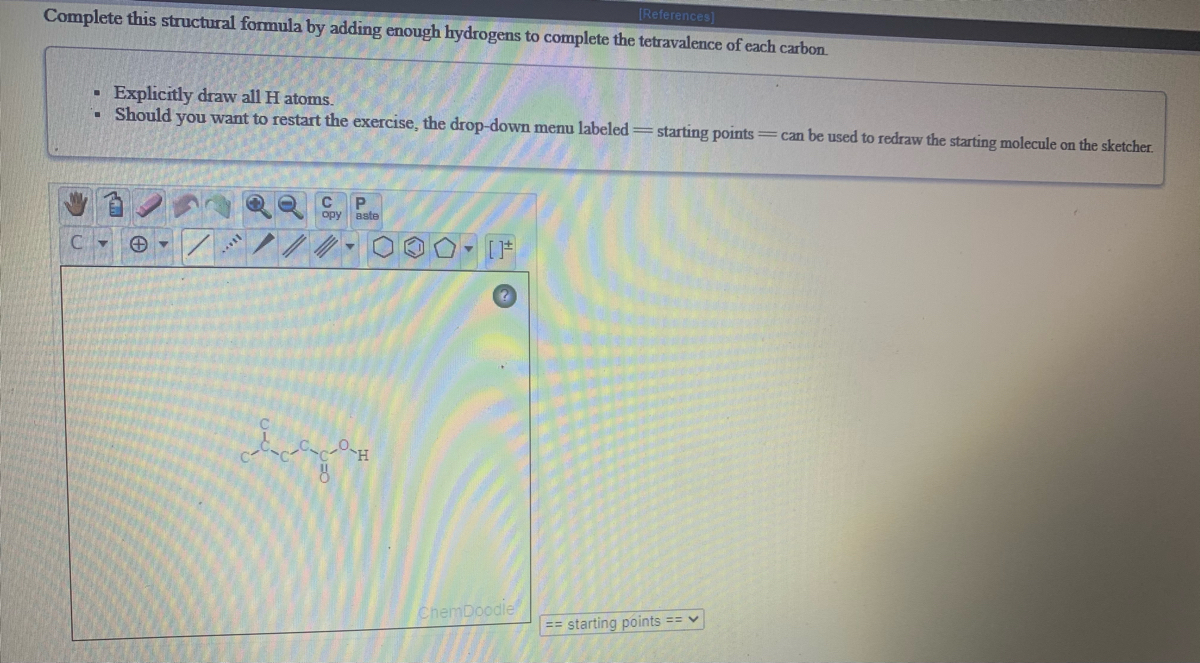
While minimizations of several discrete structures using through-space forces may look correct, please understand the theory behind the force field you are using and that any force field not specifically developed or parameterized for intermolecular forces will not lead to experimentally accurate coordinates. Please note, while you are able to optimize the entire scene, most force fields are not parameterized for multiple discrete molecular structures, and you will be relying on the through-space forces defined in the force field (mainly van der Waals and electrostatic), if defined at all. To do this, you can change the optimization scope to optimize the entire scene. RBSE, UP, MP, BIHAR BOARDQUESTION TEXT:-Write the productThe addition of hydrogen. The ChemDoodle Web Components library is a pure JavaScript chemical graphics and cheminformatics library derived from the ChemDoodle application and produced by iChemLabs. However, you may wish to optimize several discrete molecular structures at the same time and in relation to each other. P opy aate ChemDoodle Draw the product(s) of the following reactions. Hydrogens will be added for you to your structures in appropriate 3D locations. You can then simply turn on the minimizer to optimize physical coordinates. ChemDoodle 3D therefore will optimize molecular structures separately and individually as you are editing them. When building 3D structures, ChemDoodle 3D will automatically suggest the best place in three dimensions to add an atom connection. The interface was much cleaner than the free/open-source options I tried. Jmol will automatically add any files current to the state, but other files not indicated by. I never encountered a shortfall in the basic functions I required for work. No new features should be added, just bug fixes for 13.0. 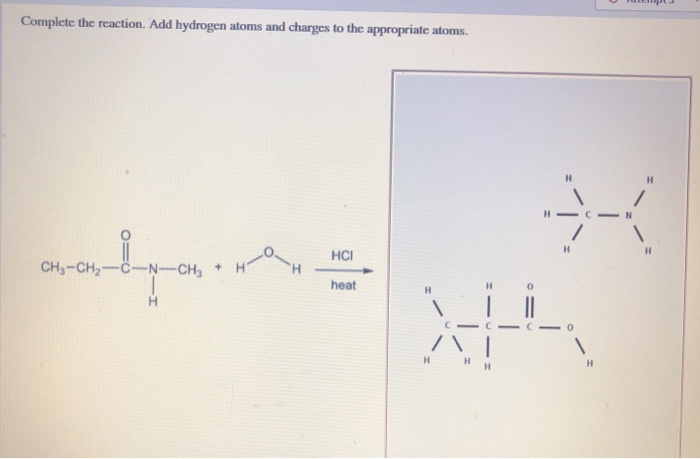
While minimizations of several discrete structures using through-space forces may look correct, please understand the theory behind the force field you are using and that any force field not specifically developed or parameterized for intermolecular forces will not lead to experimentally accurate coordinates.Most small molecule force fields are optimized for describing individual discrete molecular structures. At the time of purchase (2017) the program was a good value for a 10 pack of perpetual licenses. On this page, you can select to disable the automatic renewal of your subscription. To cancel or disable automatic renewal go to your Doodle account settings under 'subscription'. To do this, you can change the optimization scope to optimize the entire scene. Cancel your Doodle Professional subscription: A Doodle Professional subscription will be renewed automatically each year or month depending on your subscription type. However, you may wish to optimize several discrete molecular structures at the same time and in relation to each other. ChemDoodle 3D therefore will optimize molecular structures separately and individually as you are editing them. Most small molecule force fields are optimized for describing individual discrete molecular structures. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
